I am an Assistant Professor in the Leslie A. Rose Department of Mechanical Engineering at the South Dakota School of Mines and Technology. I received a Ph.D. in Civil and Environmental Engineering from Carnegie Mellon University in August 2016. After finishing my Ph.D., I joined the Department of Mathematics at Louisiana State University as a Postdoctoral Fellow and worked on numerical methods and analysis of the peridynamics theory of fracture. I then moved to the Oden Institute at UT Austin to gain experience in the computational mechanics of multiphysics and complex systems.
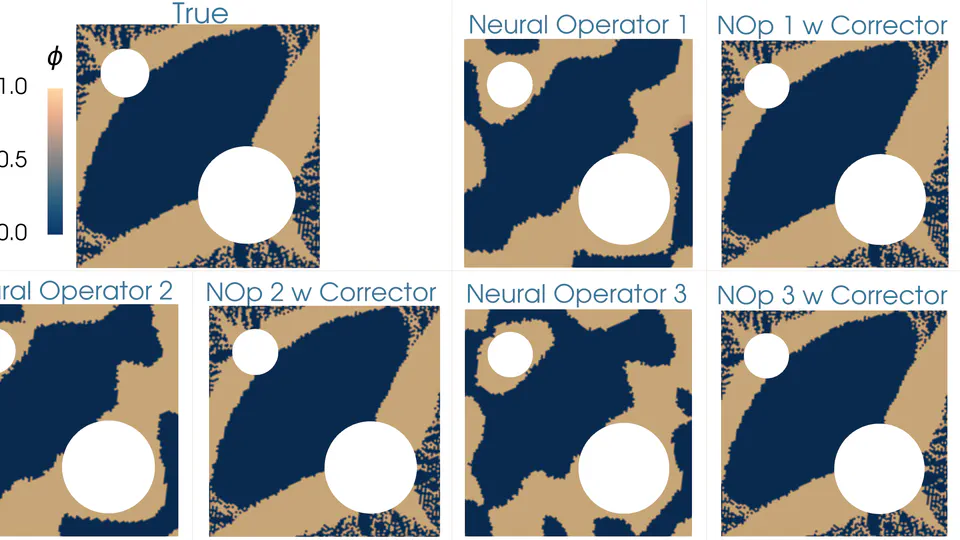
My research interests include solids and granular media mechanics, fracture mechanics, multiphysics and multiscale modeling, and applications of neural networks to engineering problems. I am serving the Journal of Peridynamics and Nonlocal Modeling as one of the associate editors, the Journal of Open Source Software (JOSS) as topic editor, and I am an Editorial Board Member of Scientific Reports.
PhD Position Available
CEAD Lab website